Flooding
Contents
Flooding#
Forecast
In this section we will produce the probabilistic rainfall assessment for a historical TC track with forecast information from JTWC issued some days prior to the TC arrival
from datetime import datetime
print("execution start: {0}".format(datetime.today().strftime('%Y-%m-%d %H:%M:%S')))
execution start: 2022-09-03 00:31:16
import os
import os.path as op
import sys
import glob
import time
from scipy.stats import qmc
import pandas as pd
import numpy as np
import xarray as xr
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
from matplotlib import gridspec
import cartopy.crs as ccrs
# raster tools
from rasterio.crs import CRS
import geopandas as gpd
import rioxarray
# bluemath
sys.path.insert(0, op.join(op.abspath(''), '..', '..'))
sys.path.insert(0, op.join(op.abspath(''), '..'))
from bluemath.teslakit.mda import MaxDiss_Simplified_NoThreshold
from bluemath.tc_warnings import download_warning
from bluemath.ddviewer.downloader.forecast import webscan_noaa_ncei, download_noaa
from bluemath.rainfall_forecast.rainfall_forecast import forecast_tracks, Reforecast_Track_Rain, \
tc_total_rain, tc_total_rain_site, Colors_basins, Plot_Rain_Hyetograph_Basins, \
Get_Analogues, Get_Flood_All_Basins, Plot_Rain_Flood_Points, \
Obtain_Basins
from bluemath.rainfall_forecast.tc_forecast import Load_TCs_Warning, Plot_tracks, Plot_Density_Forecast_Tracks
from bluemath.tc_forecast import get_info_TCs_warning, Load_TCs_Warning, Format_Tracks
from bluemath.tc_forecast import Plot_Tracks_Maxwinds, Plot_Tracks_Maxwinds_Density, Plot_Tracks_Maxwinds_zoom
# operational utils
from operational.util import clean_forecast_folder, read_config_args
Warning: ecCodes 2.21.0 or higher is recommended. You are running version 2.16.0
# aux functions
def Plot_Basins(GEO, ax=[], text=True, fill=True, figsize=[20,15]):
cols = Colors_basins(GEO)
if not ax:
fig = plt.figure(figsize=figsize)
gs = gridspec.GridSpec(1,1, wspace=0.001, hspace=0.001)
ax= plt.subplot(gs[0])
for b in np.arange(0,len(GEO)):
poly = GEO[b]
xs, ys = poly.exterior.xy
if text:
ax.text(np.double(poly.centroid.coords.xy[0][0]),np.double(poly.centroid.coords.xy[1][0]), str(b+1), fontsize=18, color='darkmagenta')#cols[b])
if fill:
ax.fill(xs, ys, alpha=0.5, fc=cols[b], linewidth=2, color=cols[b])
ax.plot(xs, ys, alpha=1, linewidth=3, color=cols[b])
ax.set_aspect('equal')
ax.set_xlabel('X UTM (m)', fontsize=20, fontweight='bold', color='navy')
ax.set_ylabel('Y_UTM (m)', fontsize=20, fontweight='bold', color='navy')
return fig
# hide warnings
import warnings
warnings.filterwarnings('ignore')
JTWC Tropical Warnings#
https://www.metoc.navy.mil/jtwc/jtwc.html
Forecast Parameters#
# project database
p_data = r'/media/administrador/HD2/SamoaTonga/data'
p_db = r'/media/administrador/HD1/DATABASES'
#site = 'Savaii' # Savaii / Upolu / Tongatapu
# (optional) get site from config file
nb_args = read_config_args(op.abspath(''), '09a_rainfall_tc_flooding')
site = nb_args['site']
print('Study site: {0}'.format(site))
Study site: Savaii
# noaa tmpfsc: automated forecast date, updated up to 3/4 days before present
years, months, days, runs = webscan_noaa_ncei()
y, m, d, h = int(years[-1]), int(months[-1]), int(days[-1]), int(runs[-1])
date_noaa = '{0:04d}{1:02d}{2:02d}'.format(y, m, d)
print('last available date for noaa tmpfsc: {0}'.format(date_noaa))
# get available run (NOAA prmsl forecast run [0, 6, 12, 18])
runs = [int(r) for r in runs]
# selected run to solve
run_sel = runs[0]
print('run: {0}'.format(run_sel))
# current date (for execution folder)
#date = datetime.today().strftime('%Y%m%d%H%M')
#print('forecast execution date: {0}'.format(date))
last available date for noaa tmpfsc: 20220829
run: 0
# forecast parameters
_clean_forecast_folder_days = 7 # days to keep after forecast folder cleaning
# rainfall footprint
run_rainfall_footprint = True
# sst data download options
_load_local_cache = False
_clean = False
# choose agencies for TC warning
ags_sel = [
' UKMI', ' JMA', ' JGSI', ' AVNI', ' HWFI', ' NVGI',
' ECM2', ' JTWC', ' JNVI', ' NVG2', ' UEMI', ' UHM2', ' UKM2',
]
# TODO REFORECAST
ags_sel=[' AVNI', ' NVGI', ' JTWC', ' NVGM',' JAVI' ]
# Hazard winds
process_raster = False
# site related parameteres
if site == 'Savaii':
# site
site_ = site.lower()
nm = 'sp'
# area of interest
area_interest = [[177, -165+360] , [-21.5, -7]]
area_interest_grid = [[187.11, 187.89],[-13.89, -13.38]]
# area_interest = [[183.826, 192.425], [-16.02, -11.6]]
# reforecast track
n, tc_name = 166, 'amos' # AMOS 2016
name_f_file = 'ash202016_amos_aids.dat'
elif site == 'Upolu':
# site
site_ = site.lower()
nm = 'up'
# area of interest
area_interest = [[177, -165+360] , [-21.5, -7]]
area_interest_grid = [[187.848, 188.63],[-14.078, -13.756]]
# reforecast track
n, tc_name = 166, 'amos' # AMOS 2016
name_f_file = 'ash202016_amos_aids.dat'
elif site == 'Tongatapu':
# site
site_ = site
nm = 'tp'
# area of interest
area_interest = [[179, -169+360] , [-23.6, -18.1]]
area_interest_grid = [[184.59, 185.041],[-21.31, -21.0275]]
# reforecast track
n, tc_name = 180, 'gita'
name_f_file = 'ash092018_gita_aids.dat'
area_interest_plot = [x for sl in area_interest for x in sl]
Database#
p_site = op.join(p_data, site)
# deliverable folder
p_deliv = op.join(p_site, 'd09_rainfall_forecast')
# Lisflood database
p_lf_db = op.join(p_db, 'Lisflood_Rainfall')
# Basins path
p_basins = op.join(p_deliv, 'basins', '{0}_db_fixed.shp'.format(nm))
# Som fitting
p_som_fit = os.path.join(p_deliv, 'som_fit_30x30_4d.nc')
# Parameters from classification
p_params_fit = os.path.join(p_deliv, 'params_fit_30x30_4d_clust.nc')
# Dem
p_dem = os.path.join(p_deliv, site_ + '_5m_dem.nc')
# LHS cases
p_lhs_subset = os.path.join(p_deliv, 'LHS_mda_sel_lisflood_volume_rain.nc')
# ibtracs coef folder
p_ibtracs_coef = op.join(p_deliv, 'TC_forecast')
# TODO REFORECAST tracks
path_tc_rf = os.path.join(p_deliv, name_f_file)
Reforecast#
date = 'reforecast_{0}'.format(tc_name)
print('forecast execution: {0}'.format(date))
forecast execution: reforecast_amos
Forecast Output Folder#
p_forecast = op.join(p_site, 'forecast', '09_rainfall_tc_inundation')
# noaa download folder
p_fore_dl_sfc = op.join(p_site, 'download_noaa', '{0}'.format(date_noaa), 'tmpsfc')
# forecast folder
p_fore_date = op.join(p_forecast, date)
print('forecast date code: {0}'.format(date))
# prepare forecast subfolders for this date
p_fore_warn = op.join(p_fore_date, 'warnings')
for p in [p_fore_date, p_fore_warn, p_fore_dl_sfc]:
if not op.isdir(p):
os.makedirs(p)
# clean forecast folder
#clean_forecast_folder(p_forecast, days=_clean_forecast_folder_days)
# clean noaa download extra folder
#clean_forecast_folder(p_fore_dl_sfc, days=_clean_forecast_folder_days)
forecast date code: reforecast_amos
# load basin data
bas_raw = gpd.read_file(p_basins)
basins = Obtain_Basins(bas_raw, site)
Download TC warnings#
# download TC warnings
#files_ash, files_bsh = download_warning(p_fore_warn, code_basin='sh')
# TODO REFORECAST
files_ash = [path_tc_rf]
files_bsh = ['test']
# stop forecast if no warnings
if len(files_bsh) == 0:
raise ValueError('No warnings in the SH. Forecast execution cancelled.')
sys.exit()
# choose storm code
code = 0
file_active = files_ash[code]
file_best = files_bsh[code]
print('active TC: {0}'.format(op.basename(file_active)))
print('best track: {0}'.format(op.basename(file_best)))
active TC: ash202016_amos_aids.dat
best track: test
# show available warning times
name_warning, available_times_warning = get_info_TCs_warning(file_active)
print('name warning: {0}'.format(name_warning))
print('last available warning time: {0}'.format(available_times_warning[-1]))
name warning: 20
last available warning time: 2016-04-25T12:00:00.000000000
# use last available time warning
time_warning = str(available_times_warning[-1])
print('time warning: {0}'.format(time_warning))
time warning: 2016-04-25T12:00:00.000000000
# TODO reforecast
name_warning = tc_name
TC Warning Output#
tc_name = str(name_warning)
p_fore_tc = op.join(p_fore_date, tc_name + '_forecast')
fore_code = 'forecast_{0}_{1}'.format(tc_name, site)
p_ref_tc = op.join(p_fore_tc, '{0}.nc'.format(fore_code))
p_ref_tc_time = op.join(p_fore_tc, '{0}_time.nc'.format(fore_code))
p_ref_tc_area = op.join(p_fore_tc, '{0}_area.nc'.format(fore_code))
p_ref_tc_area_time = op.join(p_fore_tc, '{0}_area_time.nc'.format(fore_code))
p_out_tracks = op.join(p_fore_tc, 'OUT_TRACKS')
# warnings
p_fore_save_track = op.join(p_fore_date, 'Forecast_{0}_time_{1}.nc'.format(name_warning, time_warning[:-3]))
p_fore_save_track_sel = op.join(p_fore_date, 'Forecast_{0}_time_{1}_selected.nc'.format(name_warning, time_warning[:-3]))
for p in [p_fore_tc, p_out_tracks]:
if not op.isdir(p):
os.makedirs(p)
# Load TRACKS
TRACKS = Load_TCs_Warning(file_active, time = time_warning)
# save warning TRACKS
TRACKS.to_netcdf(p_fore_save_track)
TRACKS = xr.open_dataset(p_fore_save_track)
Area#
Figure Explanation
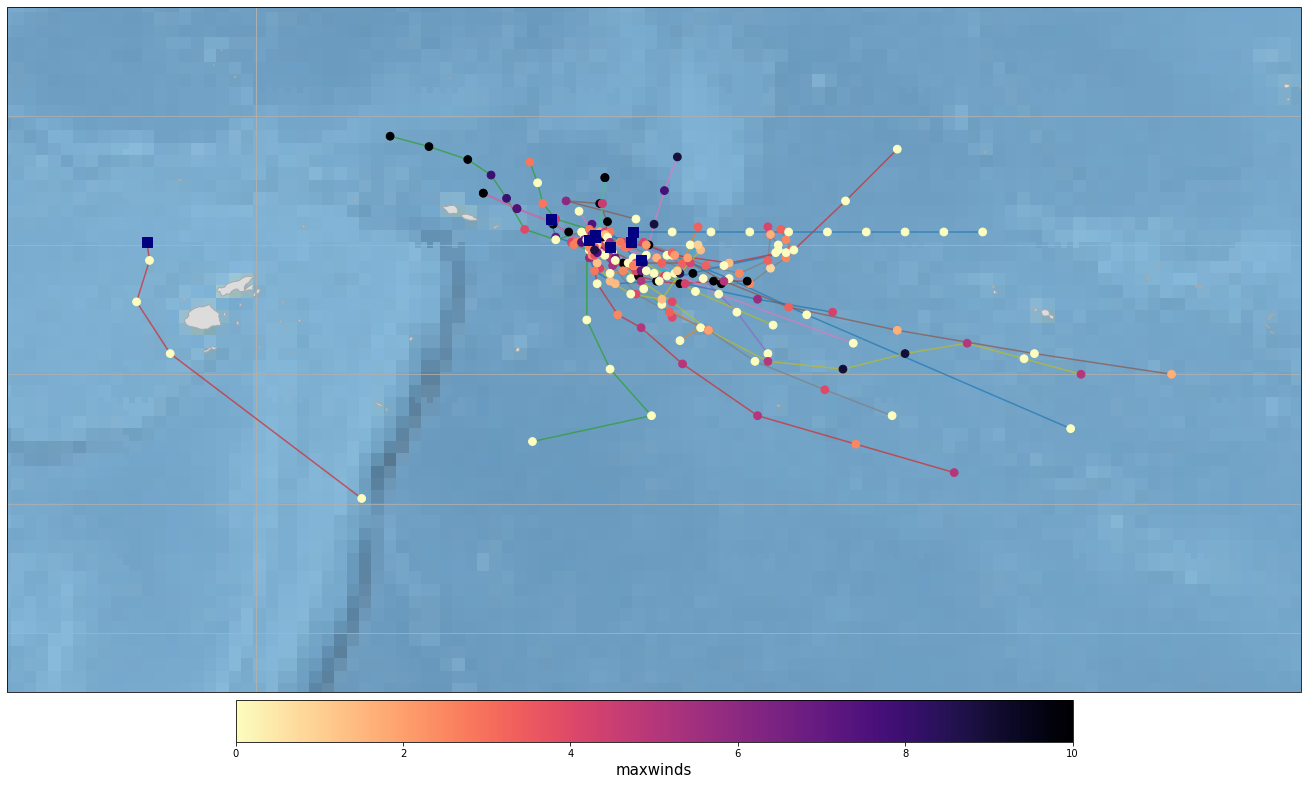
The figures below show TC warning tracks at the study site.
Need visual inspection on wether the tracks are inside the area of interest.
If there are no tracks in this area, don’t continue running the notebook.
Plot_Tracks_Maxwinds(TRACKS);
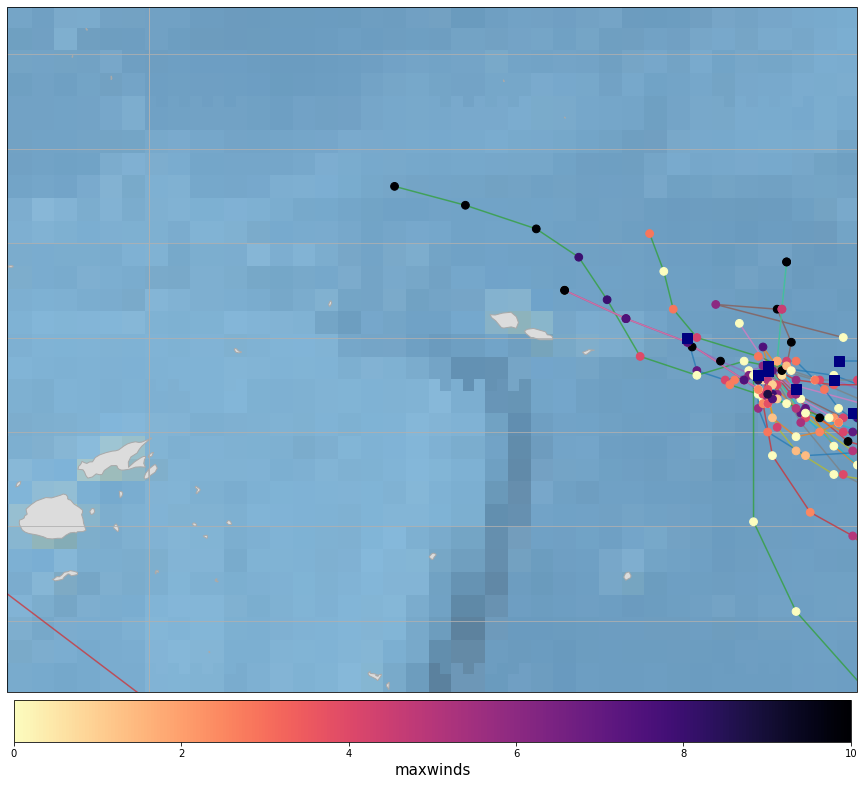
Plot_Tracks_Maxwinds(TRACKS, area_extent=area_interest_plot);
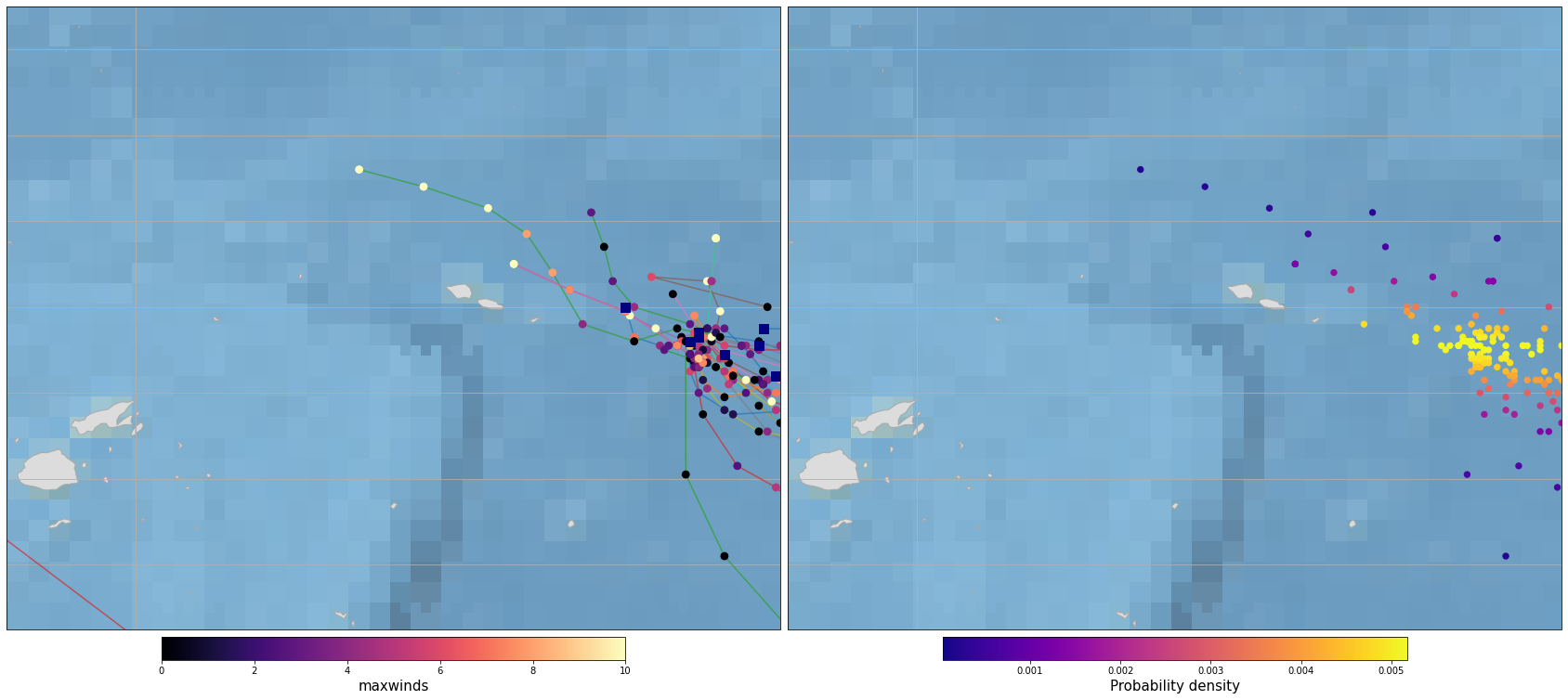
Plot_Tracks_Maxwinds_Density(TRACKS, area_extent=area_interest_plot);
Agencies selection#
print('Number of tracks: {0}'.format(len(TRACKS.track.values)))
print('Available Agencies:\n{0}\n------------'.format(TRACKS.Agency.values))
Number of tracks: 44
Available Agencies:
[' CARQ' ' AHNI' ' AVNI' ' CLIP' ' CLSW' ' CMES' ' CMSD' ' CONW' ' COT2'
' CTC2' ' DSHA' ' ECS5' ' EEM2' ' GFN2' ' JAVI' ' JJG2' ' JNVI' ' LGEA'
' LGEN' ' N012' ' N032' ' N042' ' N052' ' N062' ' N072' ' N082' ' N092'
' N102' ' N112' ' N122' ' N132' ' N142' ' N152' ' N162' ' N172' ' N202'
' NVGI' ' NVS1' ' NVS5' ' RVCN' ' S5XX' ' S5YY' ' SHIA' ' XTRP']
------------
c, ia, ib = np.intersect1d(TRACKS.Agency.values, ags_sel, return_indices=True)
print('Total number of tracks: {0}'.format(len(ia)))
print('Agencies from list available : ' + str(np.array(ags_sel)[ib]))
TRACKS = TRACKS.isel(track=ia)
TRACKS['Index'] = (('track'), ia)
Total number of tracks: 3
Agencies from list available : [' AVNI' ' JAVI' ' NVGI']
if TRACKS.track.size > 0:
Plot_Tracks_Maxwinds_Density(TRACKS, area_extent=area_interest_plot);
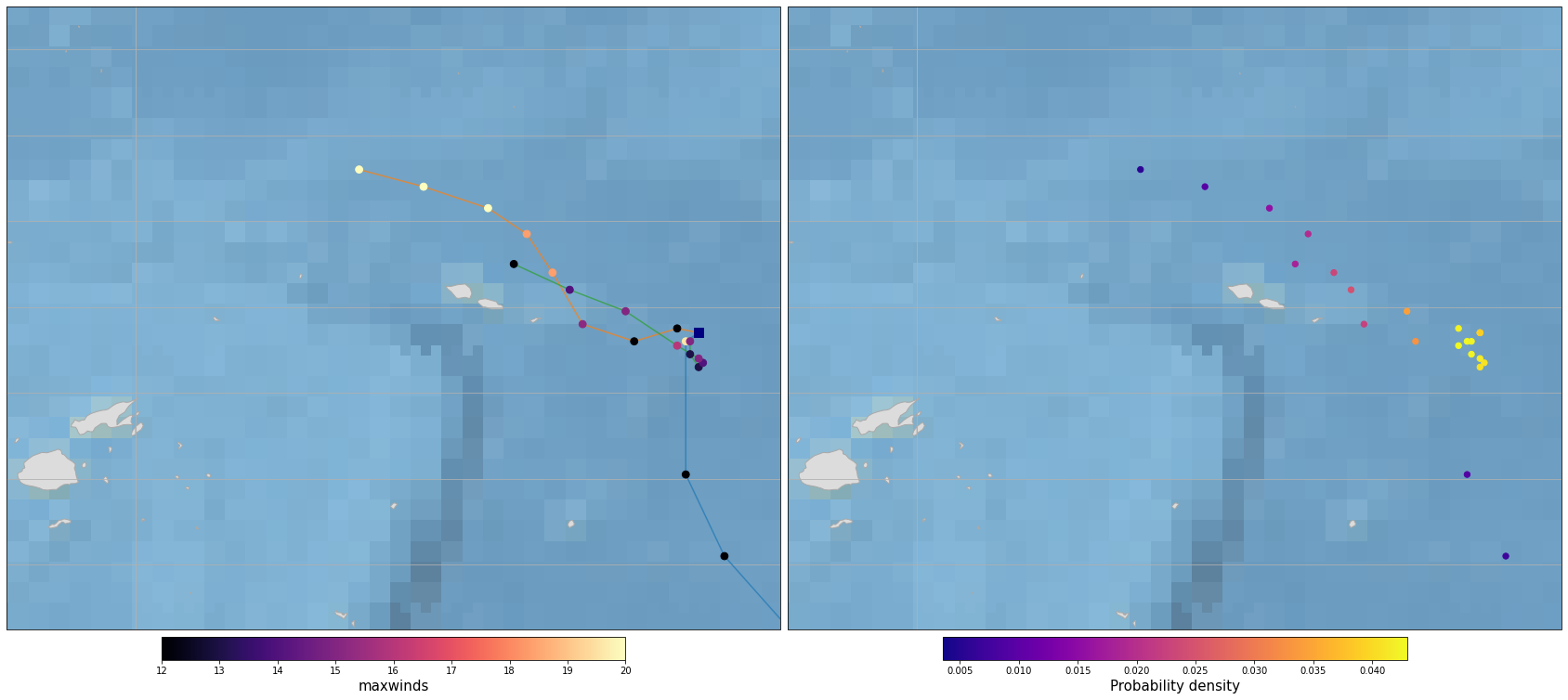
if TRACKS.track.size > 0:
Plot_Tracks_Maxwinds_zoom(TRACKS, area_interest_plot);
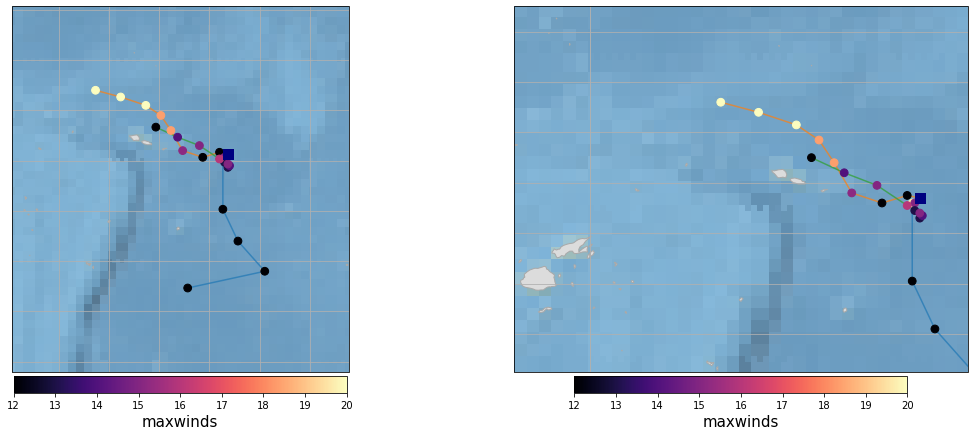
Rainfall#
Rainfall footprint#
Download needed SST data
# download SST data
#print('download tmpsfc. run number {0}'.format(run))
#sst = download_noaa(
# 'tmpsfc',
# y, m, d, h,
# p_f = p_fore_dl_sfc,
# load_local_cache = _load_local_cache, clean=_clean,
#)
#print('')
#f_save = 'tmpsfc' + '.'+str(y)+'.'+str(m)+'.'+str(hours[hi])+'.daily.grb2.nc'
#sst.to_netcdf(op.join(path_fore_dl_sfc,f_save))
# Process tracks and add sst for rainfall model
#forecast_tracks(TRACKS, sst, p_ibtracs_coef, p_fore_date, tc_name)
# TODO REFORECAST
# comentar las dos celdas de encima
Load needed files
To run the model, we need to load the data corresponding to the SOM clustering (som_fit dataset) and the parameters associated to each of the 900 clusters (params_fit dataset).
# load SOM parameteres
som_fit = xr.open_dataset(p_som_fit)
params_fit = xr.open_dataset(p_params_fit)
# run the model
if run_rainfall_footprint:
# init output holders
DSS, DSS_A = [], []
DST, DST_A = [], []
TCKS = []
for a in range(len(TRACKS.track)):
# TODO que archivo es este? el generado dentro de forecast_tracks()?
#tck = xr.open_dataset(op.join(p_fore_date, tc_name + str(a) + '.nc'))
# TODO reforecast
tck = xr.open_dataset(os.path.join(p_deliv, tc_name + '_reforecast', tc_name + str(a) + '.nc'))
# reforecast track rain
tck, dss, dst, dss_area, dst_area = Reforecast_Track_Rain(
tck, area_interest, area_interest_grid, tc_name, som_fit, params_fit,
)
DSS.append(dss)
DST.append(dst)
DSS_A.append(dss_area)
DST_A.append(dst_area)
TCKS.append(tck)
# join output
DSS = xr.concat(DSS, dim='track'); DSS.to_netcdf(p_ref_tc)
DST = xr.concat(DST, dim='track'); DST.to_netcdf(p_ref_tc_time)
DSS_A = xr.concat(DSS_A, dim='track'); DSS_A.to_netcdf(p_ref_tc_area)
DST_A = xr.concat(DST_A, dim='track'); DST_A.to_netcdf(p_ref_tc_area_time)
else:
# load reforecast track rain
DSS = xr.open_dataset(p_ref_tc)
DST = xr.open_dataset(p_ref_tc_time)
DSS_A = xr.open_dataset(p_ref_tc_area)
DST_A = xr.open_dataset(p_ref_tc_area_time)
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 121
time step 122
time step 123
time step 124
time step 125
time step 126
time step 127
time step 128
time step 129
time step 130
time step 131
time step 132
time step 133
time step 134
time step 135
time step 136
time step 137
time step 138
time step 139
time step 140
time step 141
time step 142
time step 143
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 121
time step 122
time step 123
time step 124
time step 125
time step 126
time step 127
time step 128
time step 129
time step 130
time step 131
time step 132
time step 133
time step 134
time step 135
time step 136
time step 137
time step 138
time step 139
time step 140
time step 141
time step 142
time step 143
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 121
time step 122
time step 123
time step 124
time step 125
time step 126
time step 127
time step 128
time step 129
time step 130
time step 131
time step 132
time step 133
time step 134
time step 135
time step 136
time step 137
time step 138
time step 139
time step 140
time step 141
time step 142
time step 143
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
time step 0
time step 1
time step 2
time step 3
time step 4
time step 5
time step 6
time step 7
time step 8
time step 9
time step 10
time step 11
time step 12
time step 13
time step 14
time step 15
time step 16
time step 17
time step 18
time step 19
time step 20
time step 21
time step 22
time step 23
time step 24
time step 25
time step 26
time step 27
time step 28
time step 29
time step 30
time step 31
time step 32
time step 33
time step 34
time step 35
time step 36
time step 37
time step 38
time step 39
time step 40
time step 41
time step 42
time step 43
time step 44
time step 45
time step 46
time step 47
time step 48
time step 49
time step 50
time step 51
time step 52
time step 53
time step 54
time step 55
time step 56
time step 57
time step 58
time step 59
time step 60
time step 61
time step 62
time step 63
time step 64
time step 65
time step 66
time step 67
time step 68
time step 69
time step 70
time step 71
time step 72
time step 73
time step 74
time step 75
time step 76
time step 77
time step 78
time step 79
time step 80
time step 81
time step 82
time step 83
time step 84
time step 85
time step 86
time step 87
time step 88
time step 89
time step 90
time step 91
time step 92
time step 93
time step 94
time step 95
time step 96
time step 97
time step 98
time step 99
time step 100
time step 101
time step 102
time step 103
time step 104
time step 105
time step 106
time step 107
time step 108
time step 109
time step 110
time step 111
time step 112
time step 113
time step 114
time step 115
time step 116
time step 117
time step 118
time step 119
time step 120
# TODO: reforecast
# esta celda parece que es solo para el reforecast y no debe estar en el operacional
TCKS = []
for a in range(len(TRACKS.track)):
tck = xr.open_dataset(os.path.join(p_deliv, tc_name + '_reforecast', tc_name + str(a) + '.nc'))
tck = xr.Dataset(
{
'pres_min': (['date_time'],tck.min_pres.values),
'lon': (['date_time'],tck.longitude.values),
'lat': (['date_time'],tck.latitude.values),
'sst': (['date_time'],tck.sst.values),
'time': (['date_time'],tck.time.values),
'storm_dir': (['date_time'],np.full([len(tck.sst.values)], 0)), #Concentric circles, dir doesn't matter
'sst_delta_r200': (['date_time'],tck.sst_delta.values),
'name': (tc_name)
},
coords={'date_time': tck.time.values,},
)
tck = tck.resample(date_time='1h').interpolate()
tck['time'] = (('date_time'), tck.date_time.values)
tck = tck.isel(date_time=np.where(
(tck.lon.values>area_interest[0][0]) &
(tck.lon.values<area_interest[0][1]) &
(tck.lat.values>area_interest[1][0]) &
(tck.lat.values<area_interest[1][1]))[0])
TCKS.append(tck)
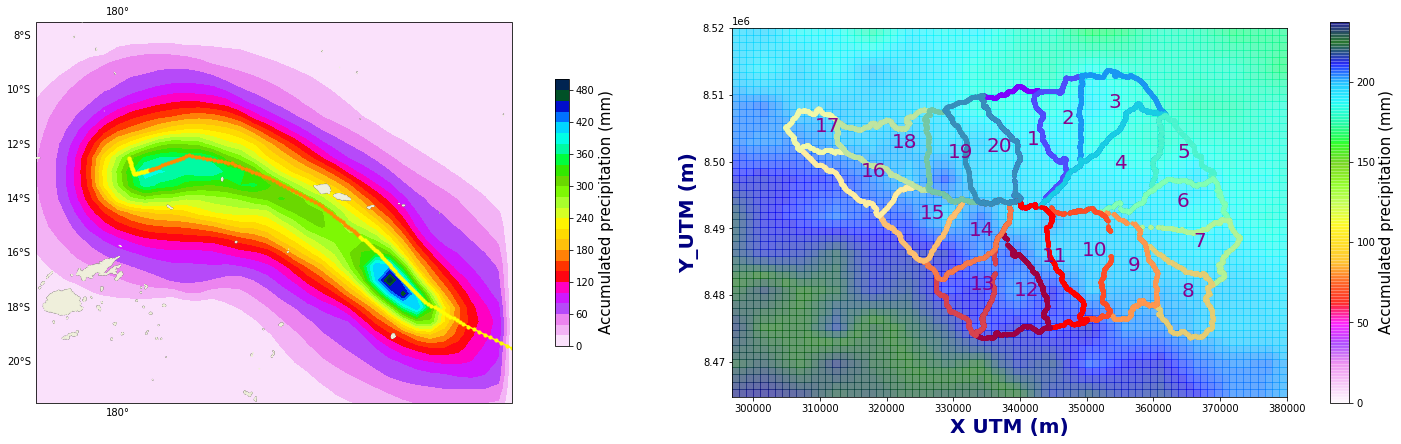
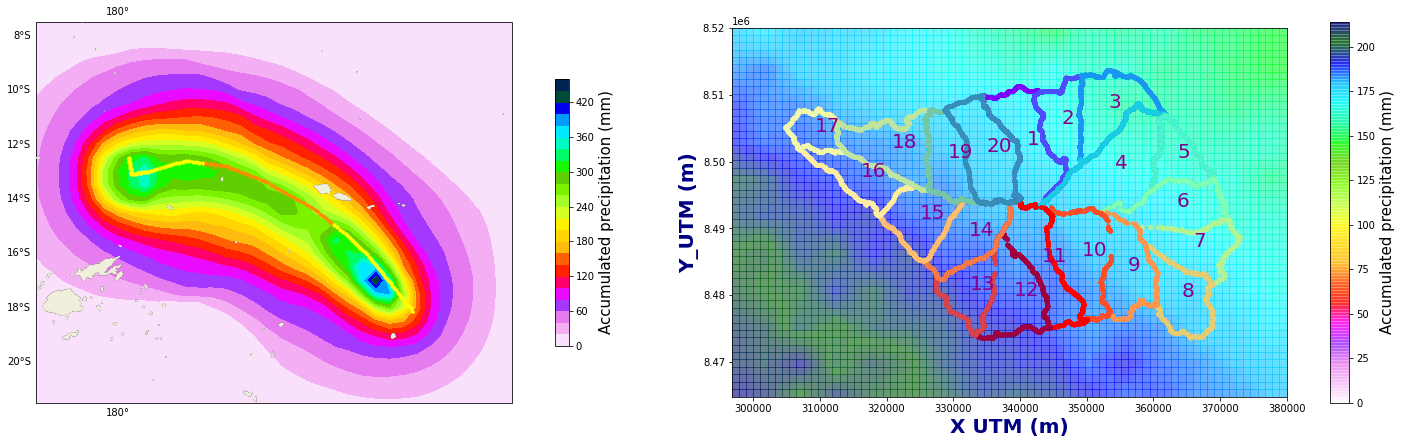
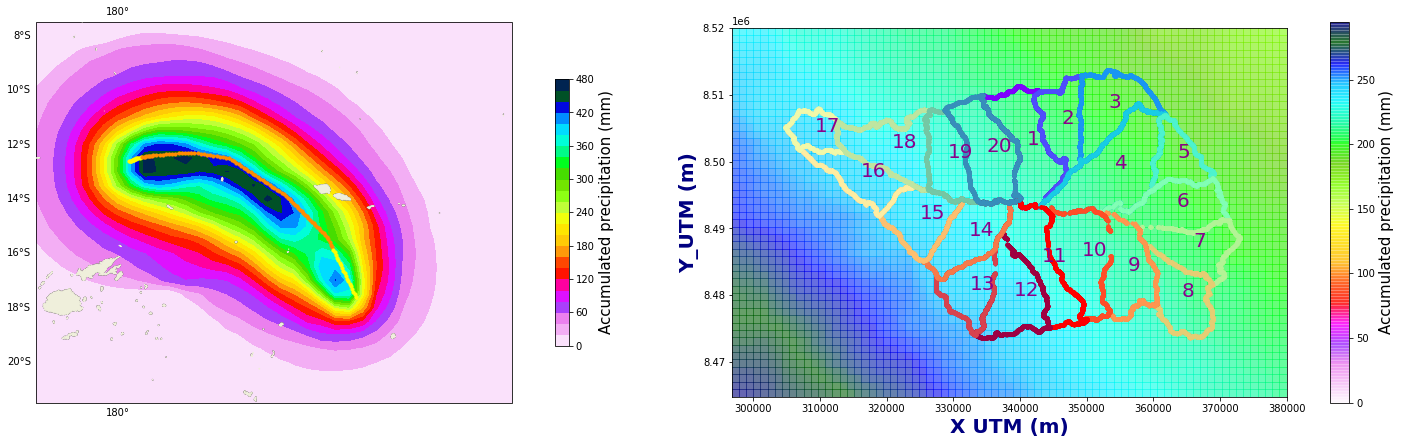
Figure Explanation
The figures below show accumulated precipitation (mm) for each track.
right panel shows a map with the basins.
# Plot each track accumulated precipitation
for t in range(len(DSS.track)):
fig = plt.figure(figsize=[25,7])
gs = gridspec.GridSpec(1, 2, wspace=0.25)
ax = fig.add_subplot(gs[0],projection = ccrs.PlateCarree(central_longitude=180))
tc_total_rain(TCKS[t], DSS.isel(track=t), fig=fig, ax=ax);
ax1=fig.add_subplot(gs[1])#,projection = ccrs.PlateCarree(central_longitude=180))
tc_total_rain_site(TCKS[t], DSS_A.isel(track=t), figsize=[25,12], basins=basins, fig=fig, ax=ax1);
SWATH#
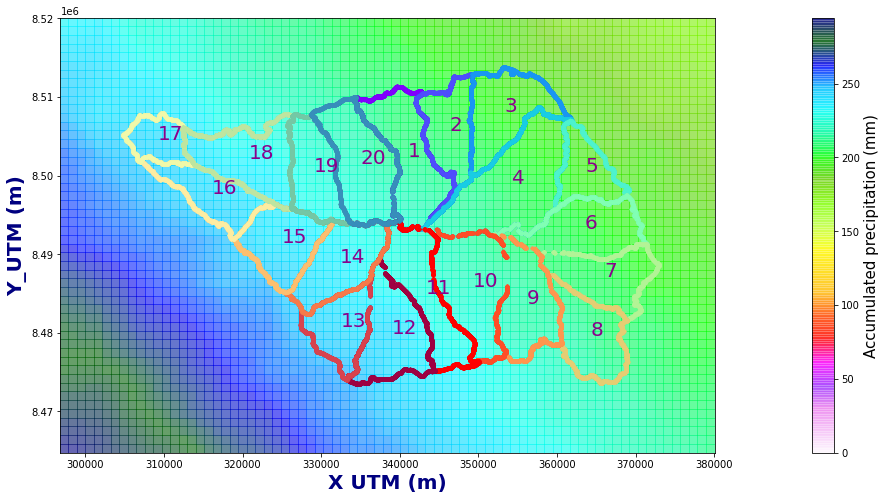
Figure Explanation
Accumualted precipitation (mm) basin maps.
fig = plt.figure(figsize=[25,8])
gs = gridspec.GridSpec(1, 1, wspace=0.25)
ax1 = fig.add_subplot(gs[0]) #,projection = ccrs.PlateCarree(central_longitude=180))
tc_total_rain_site(tck, DSS_A.max(dim='track'), figsize=[25,12], basins=basins, fig=fig, ax=ax1);

Probabilistic assessment#
lims_dss = [150, 200, 250, 300]
for l in lims_dss:
DSS['rain_prob'+str(l)+'mm'] = (('lat', 'lon'), np.where(DSS.rain>l,1,0).sum(axis=(0))/(len(DSS.track.values)))
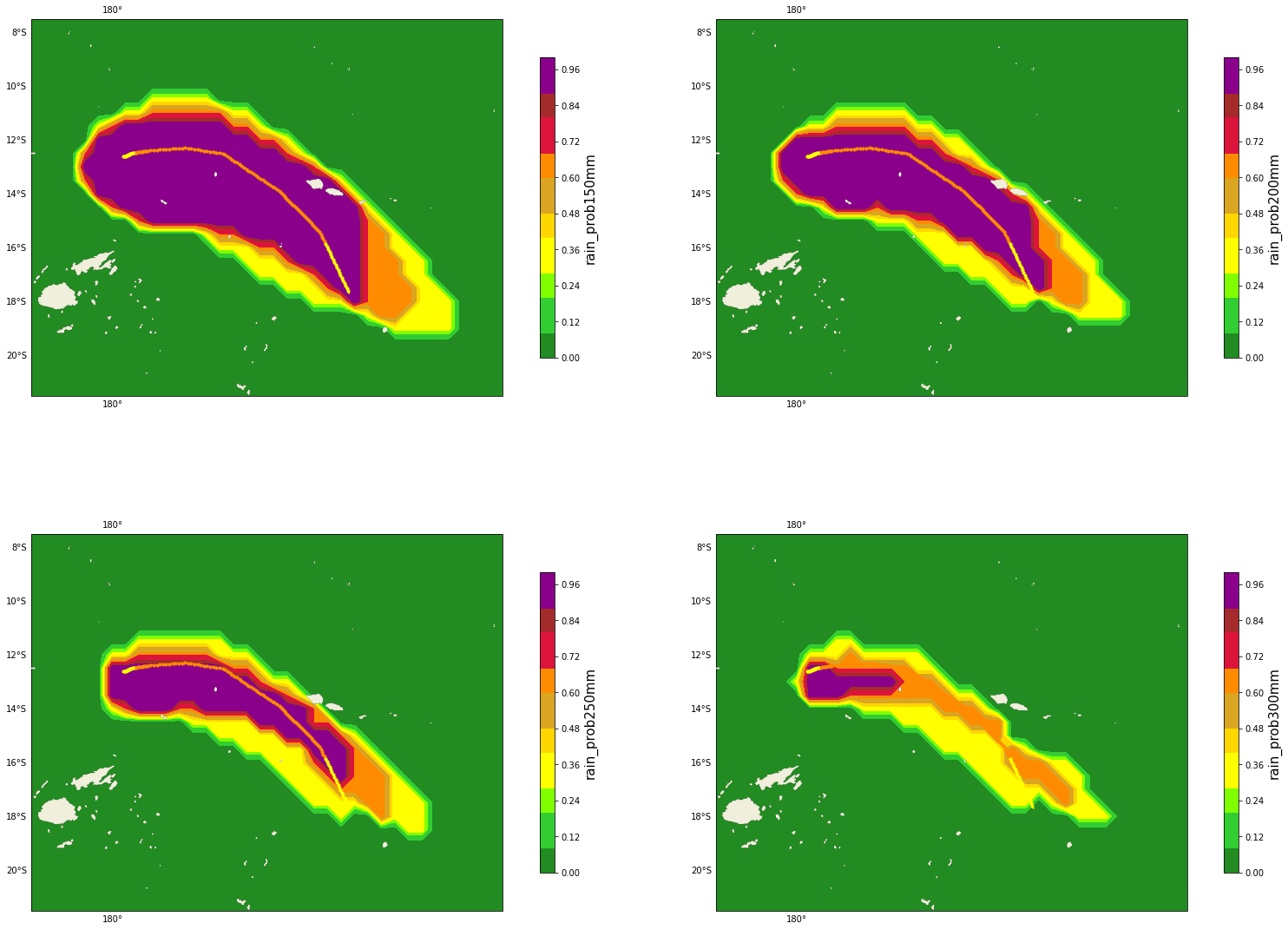
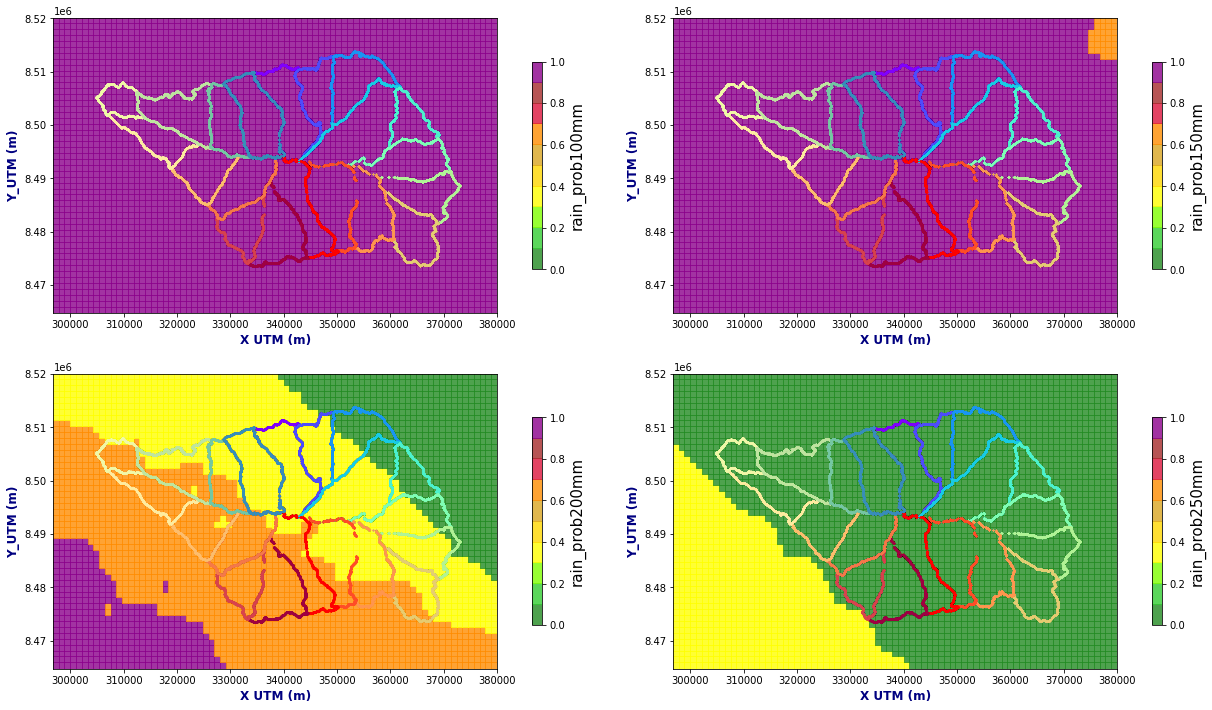
Figure Explanation
Rainfall higher than threshold probability maps.
fig = plt.figure(figsize=[25, 20])
gs = gridspec.GridSpec(2,2, wspace=0.3)
for a in range(len(lims_dss)):
ax = fig.add_subplot(gs[a], projection = ccrs.PlateCarree(central_longitude=180))
tc_total_rain(
TCKS[t], DSS.isel(track=t),
var='rain_prob'+str(lims_dss[a])+'mm',
fig=fig, ax=ax,
);
lims_dssa = [100, 150, 200, 250]
for l in lims_dssa:
DSS_A['rain_prob'+str(l)+'mm'] = (('lat', 'lon'), np.where(DSS_A.rain>l,1,0).sum(axis=(0))/(len(DSS_A.track.values)))
fig = plt.figure(figsize=[20, 12])
gs = gridspec.GridSpec(2,2, wspace=0.25)
for a in range(len(lims_dssa)):
ax1 = fig.add_subplot(gs[a])
tc_total_rain_site(
tck, DSS_A, font_s=12,mk_s=1, text=False,
var='rain_prob'+str(lims_dssa[a])+'mm', basins=basins,
fig=fig, ax=ax1,
);
Flooding#
Flooding footprint#
# lhs subset
n_cases = 50
df_v = xr.open_dataset(p_lhs_subset).isel(index=range(n_cases))
subset_norm = np.vstack([
(df_v['V'].values-np.nanmean(df_v['V'].values))/np.std(df_v['V'].values),
(df_v['I*'].values-np.nanmean(df_v['I*'].values))/np.std(df_v['I*'].values),
]).T
# dem
dem = xr.open_dataset(p_dem)
if 'z' in dem.data_vars:
dem = dem.rename({'z':'elevation'})
res_f = 2
dem = dem.isel(x=np.arange(0, len(dem.x.values),res_f), y = np.arange(0, len(dem.y.values),res_f))
dem['elevation'] = (('y', 'x'), np.where(dem.elevation.values < 0, np.nan, dem.elevation.values))
Figure Explanation
This figure show the basins maps.
# plot area basins
Plot_Basins(basins, figsize = [20, 15]);
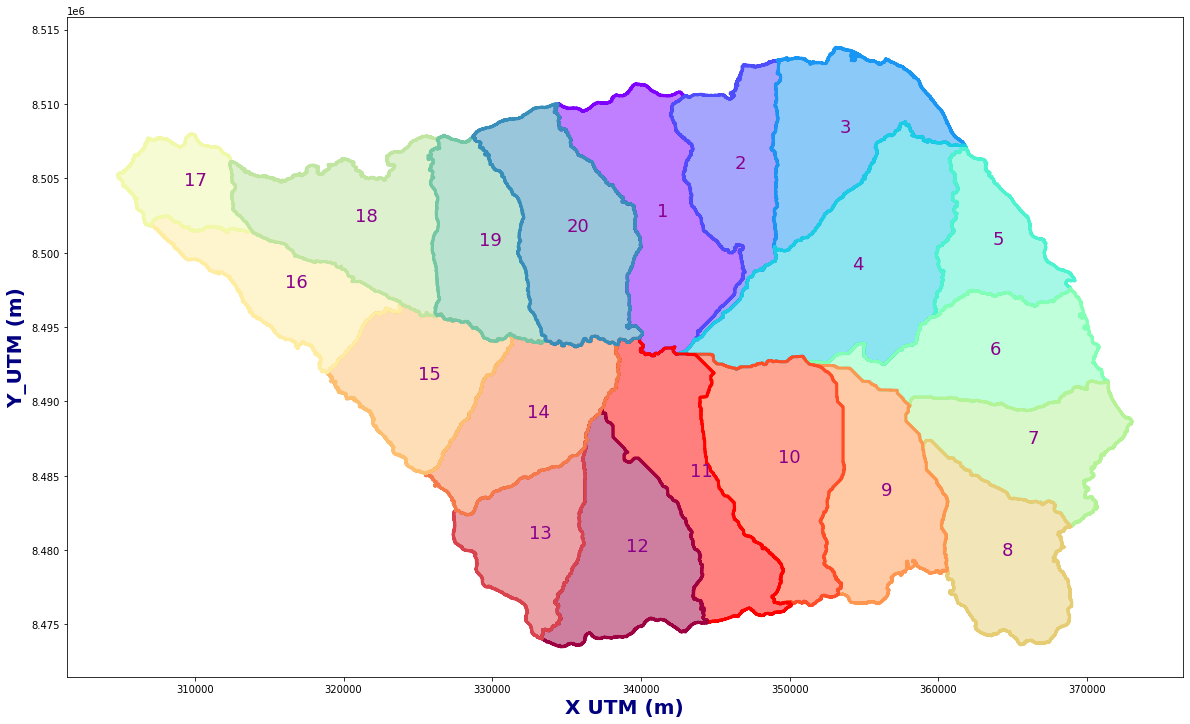
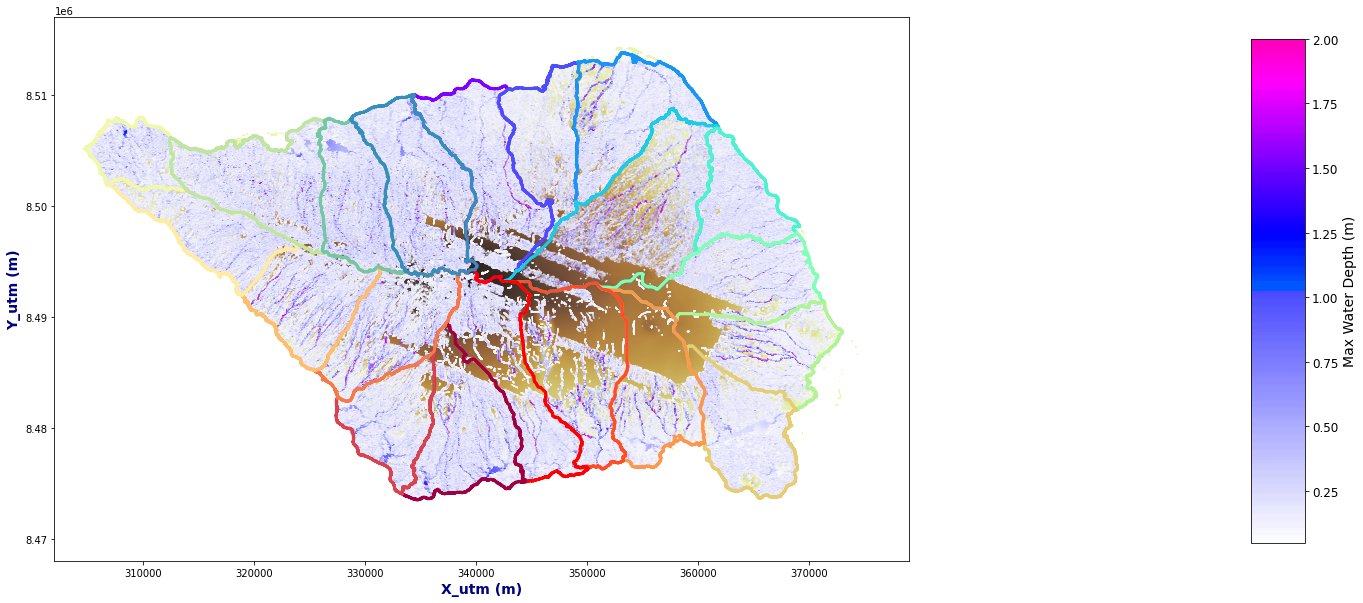
Figure Explanation
TODO: FIGURE EXPLANATION.
# iterate each track
for tk in range(len(DSS.track)):
print('track : ' + str(tk))
dst_area = DST_A.isel(track=tk)
dss_area = DSS_A.isel(track=tk)
# Calculate mean rain per basin
x_utm, y_utm, INFO = tc_total_rain_site(
TCKS[tk], dss_area,
figsize=[25,10], rain=0,
figp=False,
basins=basins, points=1,
#vmax=970,
)
dss_area['X_UTM'] = (('lon'), x_utm)
dss_area['Y_UTM'] = (('lat'), y_utm)
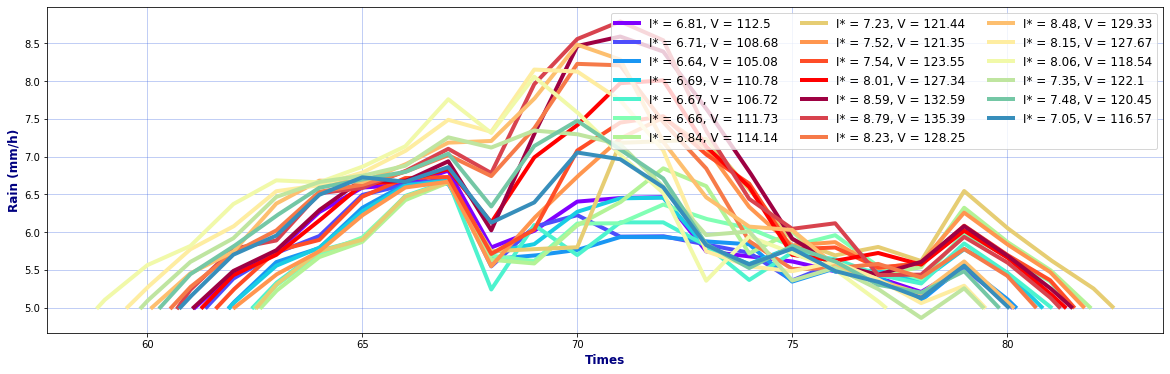
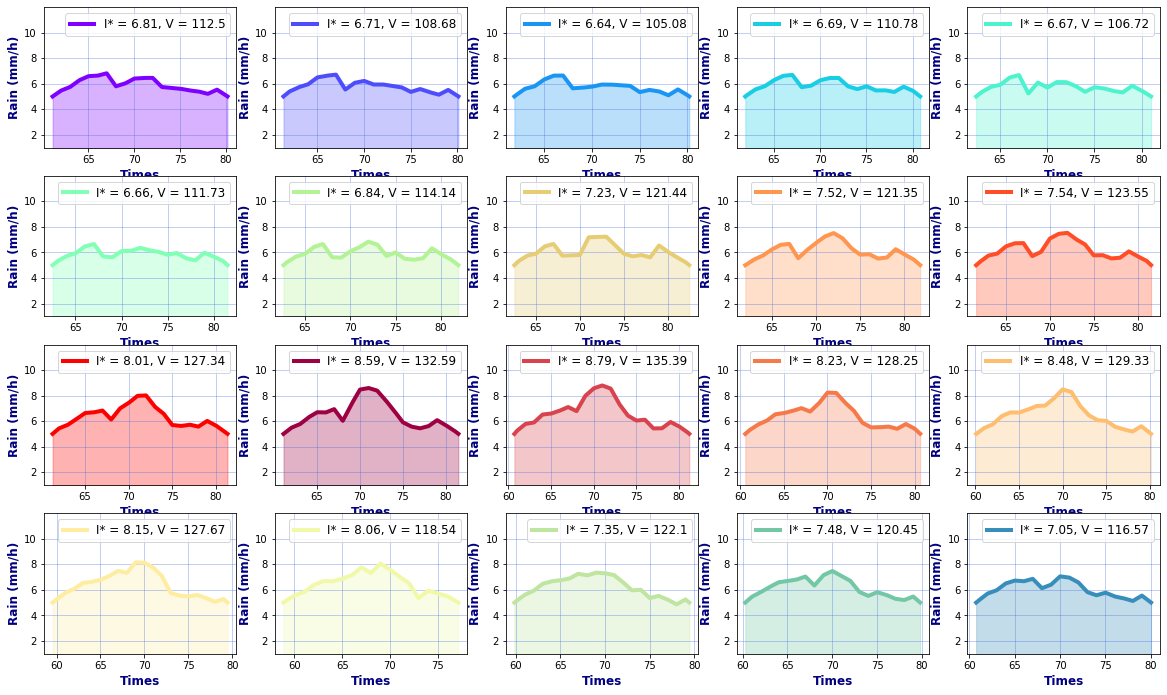
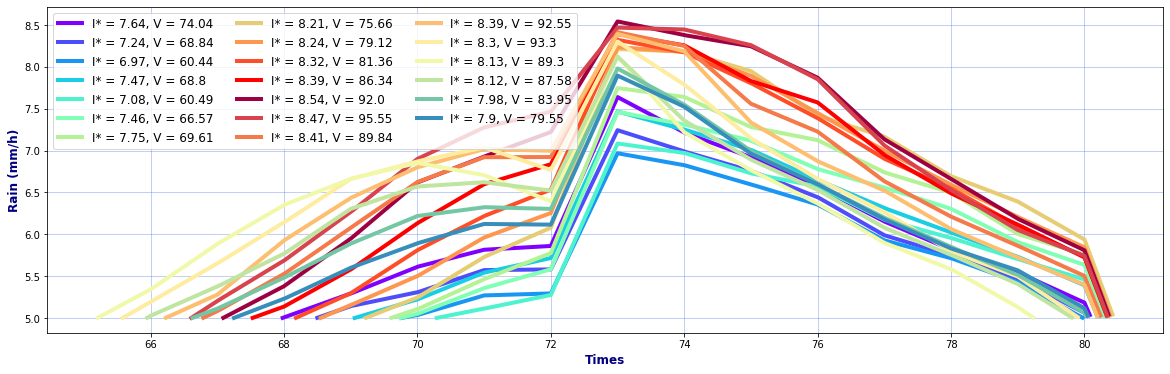
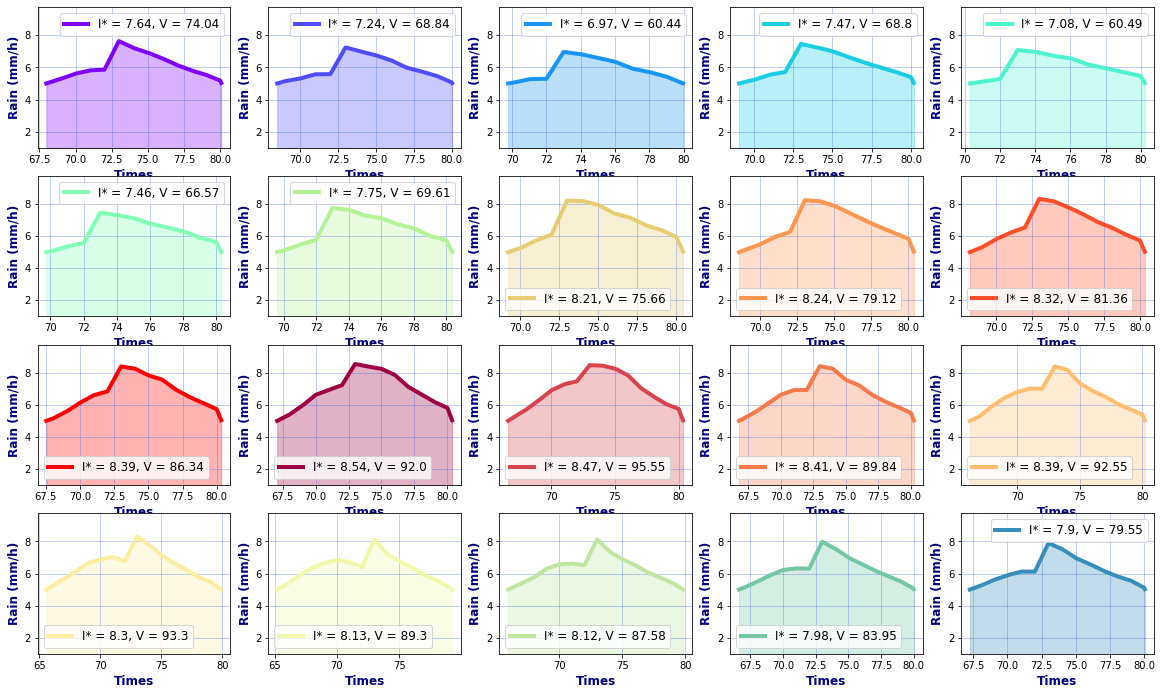
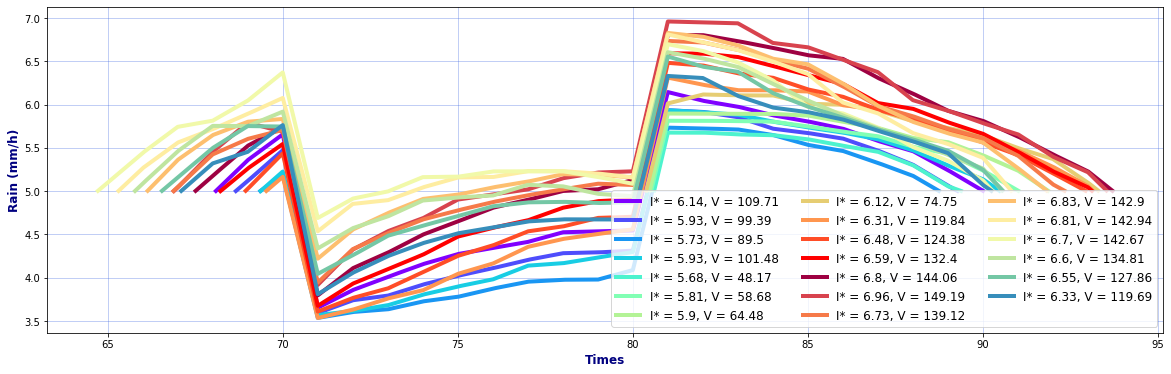
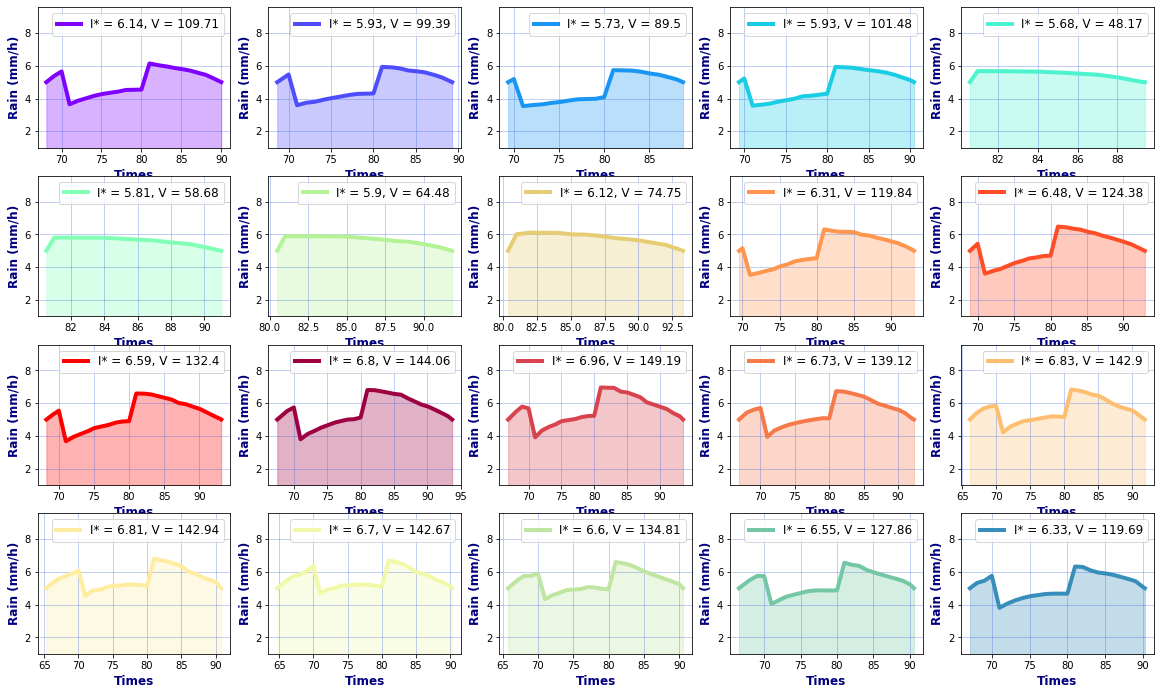
# Hyetographs per basin
rain, _, _ = Plot_Rain_Hyetograph_Basins(
dst_area, dss_area,
basins, INFO, site,
deltat=1, thr_plot=1, thr=5,
fgsize1 = [20,6], fgsize2 = [20,15],
)
# Normalize rain to find analogues
tc_rain_norm = np.vstack([
(rain['Volume'].values-np.nanmean(df_v['V'].values))/np.std(df_v['V'].values),
(rain['I'].values-np.nanmean(df_v['I*'].values))/np.std(df_v['I*'].values),
]).T
# Analogue calculations
n_an = 3 # Number of analogue points
AN = Get_Analogues(n_an, tc_rain_norm, subset_norm, df_v)
# TODO
# Plot_Analogues(df_v, rain, AN, basins, figsize=[12,12])
OUT = Get_Flood_All_Basins(basins, AN, p_lf_db, nm)
# store track output
np.savetxt(op.join(p_out_tracks, 'Flooding_Metamodel_Analogues_{0}_{1}_xyz_points.nc'.format(tc_name, tk)), OUT.values)
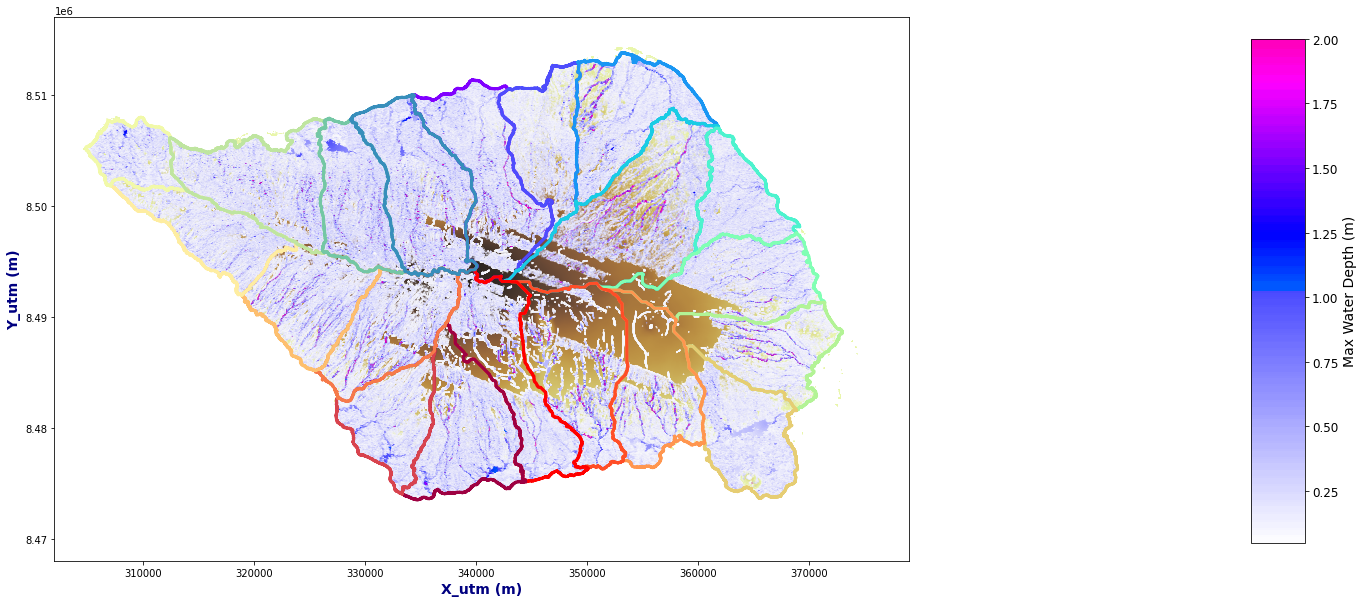
extent = [dem.x.min()-1000, dem.x.max()+1000, dem.y.min()-1000, dem.y.max()+1000]
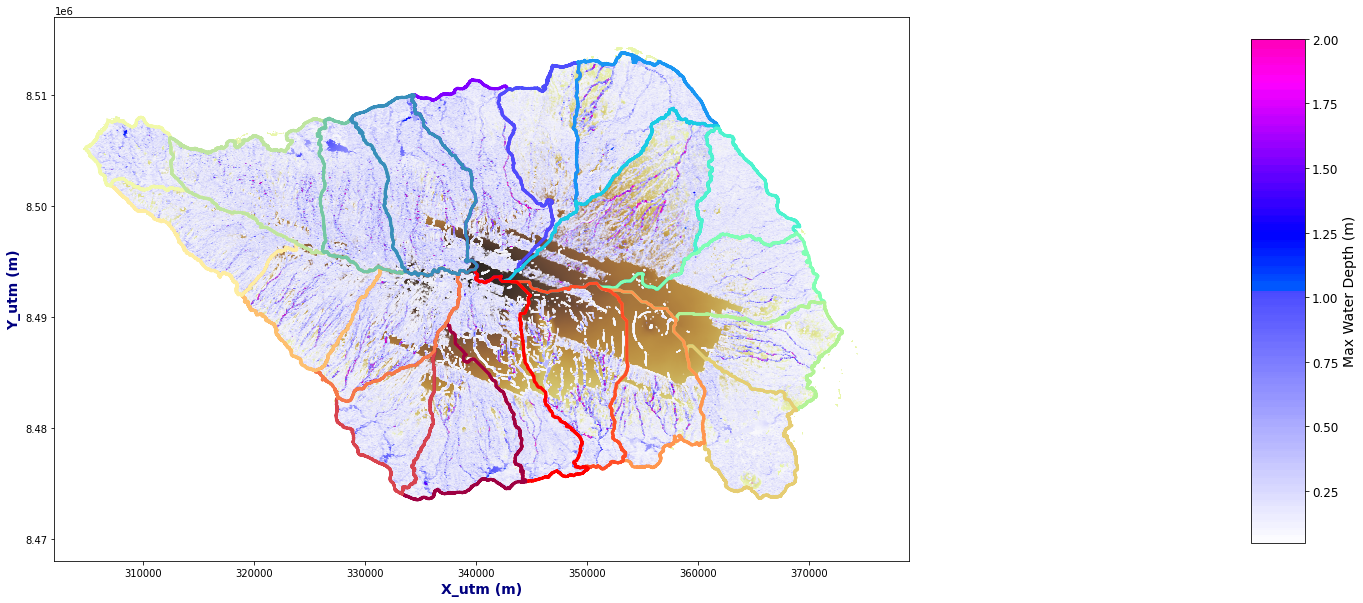
Plot_Rain_Flood_Points(basins, OUT, dem=dem, extent=extent)
if tk==0:
OUT_C = OUT.dropna().set_index(['x','y'])
else:
OUT_C = pd.concat([OUT_C, OUT.dropna().set_index(['x','y'])], axis=0, join='outer')
OUT_C = OUT_C.groupby(['x', 'y'])['z'].max().to_frame()
OUT_C = OUT_C.reset_index()
# store env output
OUT_C.to_xarray().rename({'index':'point'}).to_netcdf(op.join(p_out_tracks, 'Flooding_Metamodel_Analogues_envelope.nc'))
track : 0
Basin 1
Basin 2
Basin 3
Basin 4
Basin 5
Basin 6
Basin 7
Basin 8
Basin 9
Basin 10
Basin 11
Basin 12
Basin 13
Basin 14
Basin 15
Basin 16
Basin 17
Basin 18
Basin 19
Basin 20
track : 1
Basin 1
Basin 2
Basin 3
Basin 4
Basin 5
Basin 6
Basin 7
Basin 8
Basin 9
Basin 10
Basin 11
Basin 12
Basin 13
Basin 14
Basin 15
Basin 16
Basin 17
Basin 18
Basin 19
Basin 20
track : 2
Basin 1
Basin 2
Basin 3
Basin 4
Basin 5
Basin 6
Basin 7
Basin 8
Basin 9
Basin 10
Basin 11
Basin 12
Basin 13
Basin 14
Basin 15
Basin 16
Basin 17
Basin 18
Basin 19
Basin 20
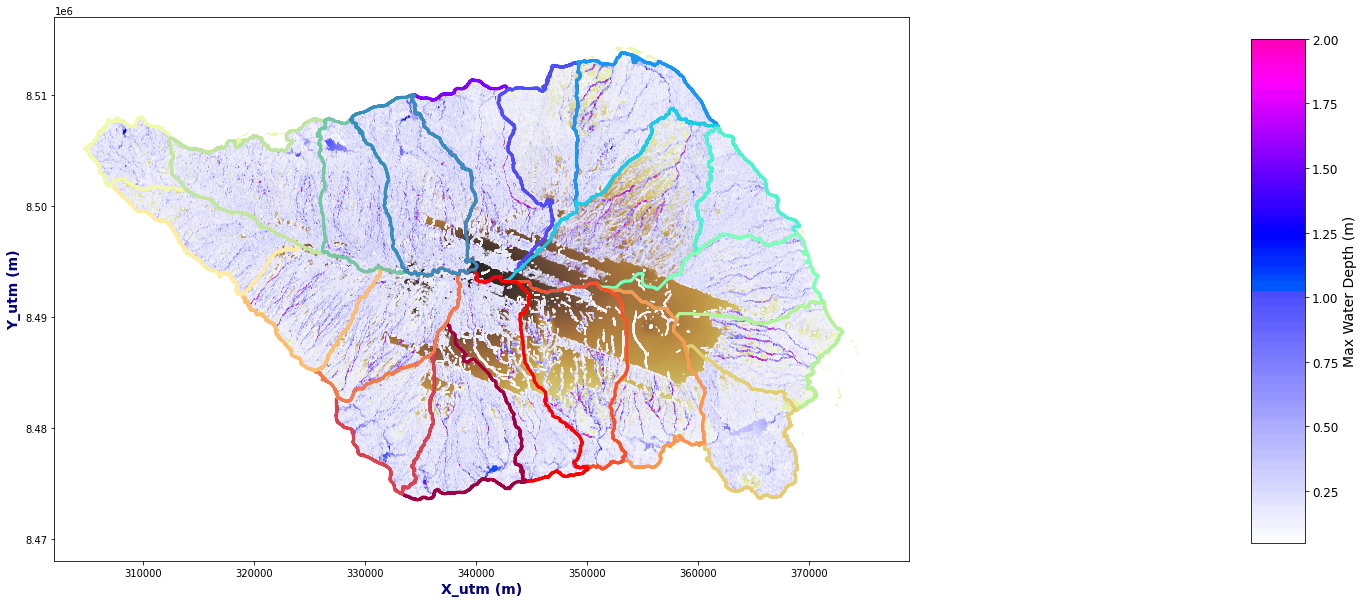
SWATH#
Figure Explanation
TODO: FIGURE EXPLANATION.
Plot_Rain_Flood_Points(basins, OUT_C, dem=dem, extent=extent);
print("execution end: {0}".format(datetime.today().strftime('%Y-%m-%d %H:%M:%S')))
execution end: 2022-09-03 01:06:42

